Section:
New Results
A probabilistic framework to infer brain functional connectivity from anatomical connections
We present a novel probabilistic framework to learn across several
subjects a mapping from brain anatomical connectivity to functional
connectivity, i.e. the covariance structure of brain activity. This
prediction problem must be formulated as a structured-output learning
task, as the predicted parameters are strongly correlated. We
introduce a model selection framework based on cross-validation with
a parametrization-independent loss function suitable to the manifold
of covariance matrices. Our model is based on constraining the
conditional independence structure of functional activity by the
anatomical connectivity. Subsequently, we learn a linear predictor of
a stationary multivariate autoregressive model. This natural
parameterization of functional connectivity also enforces the
positive-definiteness of the predicted covariance and thus matches
the structure of the output space. Our results show that functional
connectivity can be explained by anatomical connectivity on a
rigorous statistical basis, and that a proper model of functional
connectivity is essential to assess this link.
See also [20] and Fig. 5 .
Figure
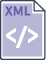
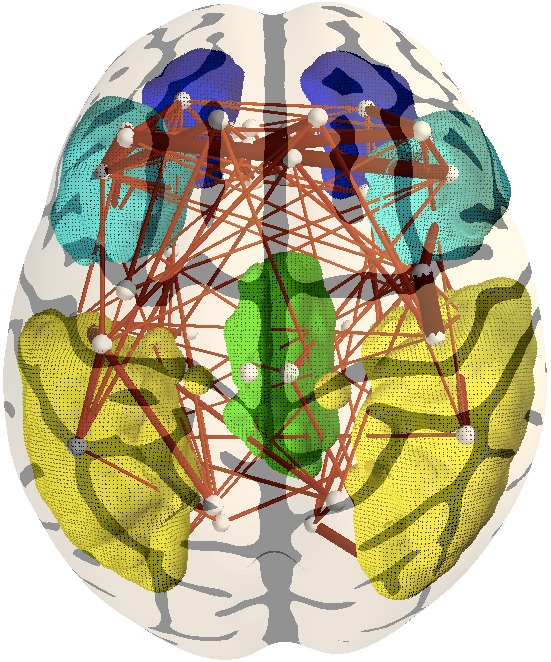
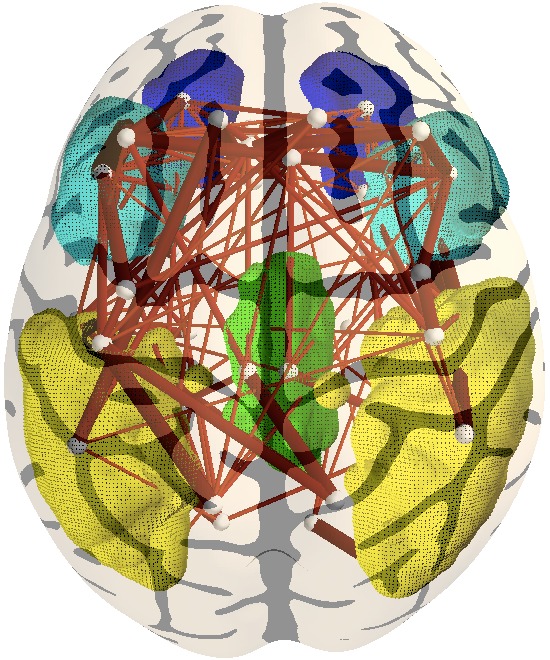
5. Identifying structural connections associated with the
default mode network. With yellow is represented the lateral
parietal cortex, green areas represent the posterior cingulate gyrus
(PCC ), blue and light blue represent the medial prefrontal and
orbito-frontal areas, respectively. The right model performs much
better in terms of cross-validated data likelihood. |